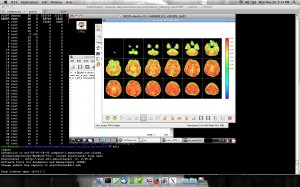
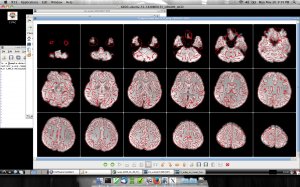
The Human Connectome Project aims to provide an unparalleled compilation of neural data, an interface to graphically navigate this data and the opportunity to achieve never before realized conclusions about the living human brain.
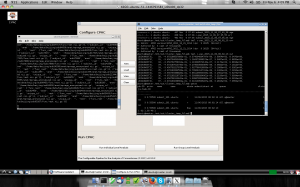
The Configurable Pipeline for the Analysis of Connectomes (C-PAC) is an open-source software pipeline for automated preprocessing and analysis of resting-state fMRI data. C-PAC builds upon a robust set of existing software packages including AFNI, FSL, and ANTS, and makes it easy for both novice users and experts to explore their data using a wide array of analytic tools. Users define analysis pipelines by specifying a combination of preprocessing options and analyses to be run on an arbitrary number of subjects. Results can then be compared across groups using the integrated group statistics feature.
StarCluster is an open source cluster-computing toolkit for Amazon’s Elastic Compute Cloud (EC2).
StarCluster allows anyone to easily create a cluster computing environment in the cloud suited for distributed and parallel computing applications and systems. It is designed to automate and simplify the process of building, configuring, and managing clusters of virtual machines on Amazon’s EC2 cloud.
Some features of StarCluster includes:
- Clusters are automatically configured with NFS, Open Grid Scheduler (formerly SGE) queuing system
- Support for attaching and NFS-sharing Amazon Elastic Block Storage (EBS) volumes for persistent storage across a cluster
- Comes with publicly available Ubuntu-based Amazon Machine Images (AMI) configured for distributed and parallel computing.
- AMI includes OpenMPI, OpenBLAS, Lapack, NumPy, SciPy, and other useful scientific libraries.
- Ability to Add/Remove Nodes
After attending the Python Philippines Conference 2016 last February, we were fascinated by speakers dealing with big data and processing pipelines.
I would like to share these tool sets which make use of Amazon EC2 in running jobs using clusters to process pipelines.
We got a chance to work with projects “Human Connectome” and we were using CPAC for processing datasets of Functional Magnetic Resonance Imaging (fMRI).
In our project the basic concept is to configure your StarCluster to run CPAC AMI, specify instance types, number of nodes of a cluster where these cluster is running Open Grid Engine (formerly Sun Grid Engine) for queuing jobs. The results of each jobs are copied to a shared directory or can be uploaded to a specific S3 buckets.
Here are some screenshots we have running in customized CPAC AMI:
In cases where researchers, students and enthusiasts who wants to process scientific pipelines in the cloud using Amazon EC2 and using these toolset ex. Starcluster (in creating clusters) can be a good way to speed up the processing. The nice value is we can choose high memory/cpu for these instances and it is pay per use. Which would save as more time and money than acquiring them on our own or for our premises.
I’m not sure with this idea, but I am proposing to have a web dashboard where we can define our instance types, pipelines and cluster configurations from a web dashboard and graphically run these processing inside EC2. Output of the processing can be download via S3 or uploaded to another server. Unfortunately StarCluster can only run inside Amazon EC2, AMI must be baked before it can be used, or if we want to customize we need to use Starcluster plugins.
Hopefully it could bring another tool like ElasticCluster into the scene. Where it can be run on any cloud provider like Openstack and other cloud platforms, and has simple configuration file to define cluster template.
Refences:
http://star.mit.edu/cluster/
http://fcp-indi.github.io/
http://www.humanconnectomeproject.org/